Time: 05:00 PM – Amsterdam, Berlin, Rome, Stockholm, Vienna
Date: March 30, 2023
Topics/Speakers:

Therapeutic application of engineered U1 snRNAs in retinopathies and other disorders
Paolo Pigini

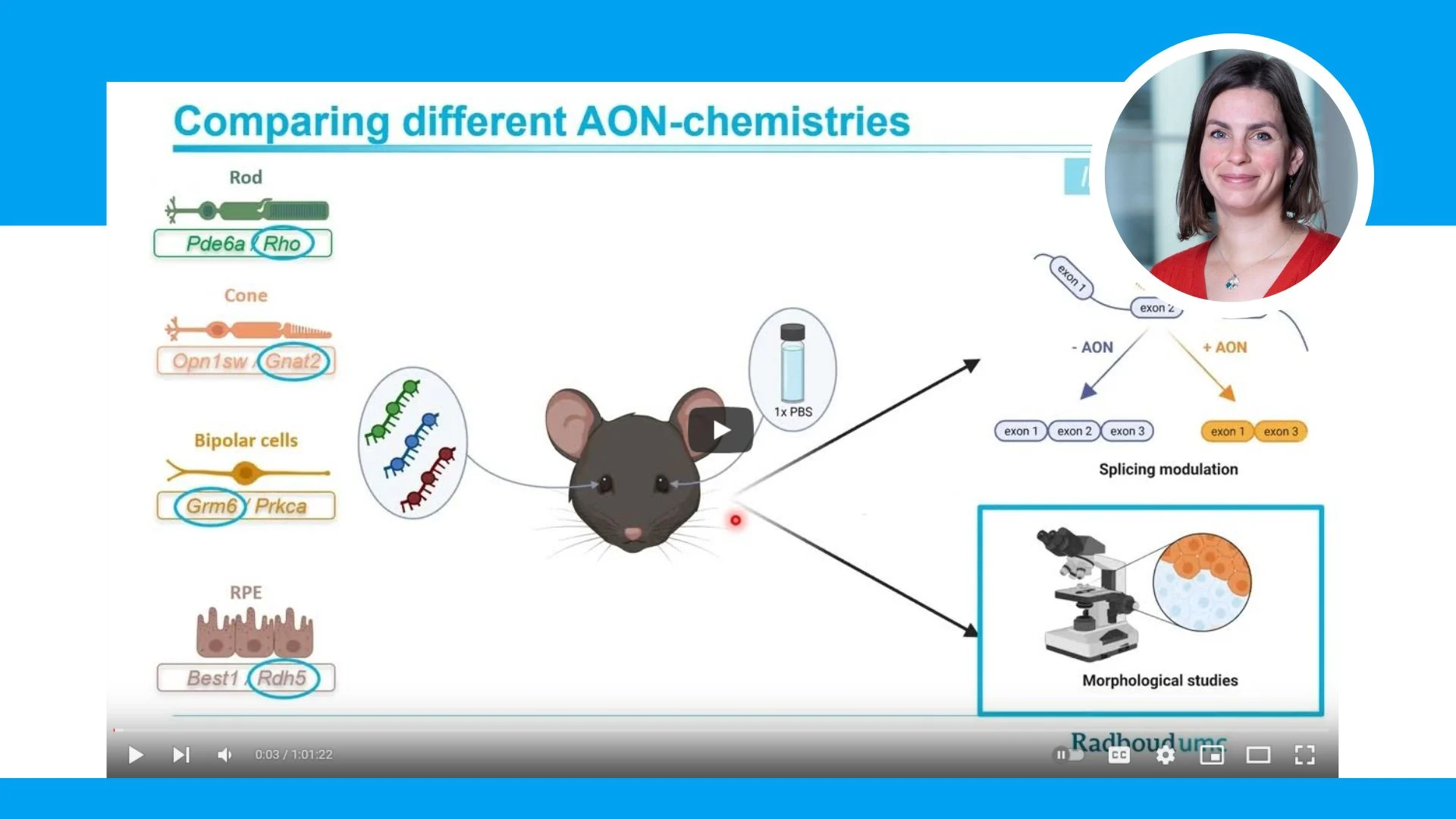
Assessing efficacy and safety of delivery of different chemically modified antisense oligonucleotides to the eye
Irene Vazquez Dominguez
Irene will present an overview of the research including the design of the ASOs and midigenes we employed in the in vitro studies as well as the considerations for in vivo delivery of the chemically modified ASOs. In addition, she will highlight the most important results obtained in this research either in vitro (RT-PCR studies) or in vivo (RT-PCR studies, morphological studies, etc).






