Looking for something?
Could a Single Oligonucleotide Medication Treat Multiple Different Diseases?

Our relatively new field is swiftly growing, and another exciting novel therapeutic RNA modality is now gaining traction. tRNA is vital in protein synthesis and by engineering tRNA molecules, it may be possible to treat disease. In fact, it is thought that a single tRNA therapy could have the potential to treat several different diseases.
The Essential Function of tRNA
Much work has been done on understanding the role of mRNA in the disease process and on understanding and developing antisense oligonucleotides (ASOs) and siRNA therapeutics to treat disease by interacting with mRNA. It is well understood that as mRNA is translated into protein, mutations can cause disruptions to this process, including the production of mutated, toxic proteins, or too much or too little of a protein being created, resulting in diseases that are often debilitating and deadly.
However, there is another small RNA molecule that is vital to protein synthesis called transfer RNA (tRNA). Each tRNA molecule contains an anticodon region and a region that attaches to a specific amino acid.
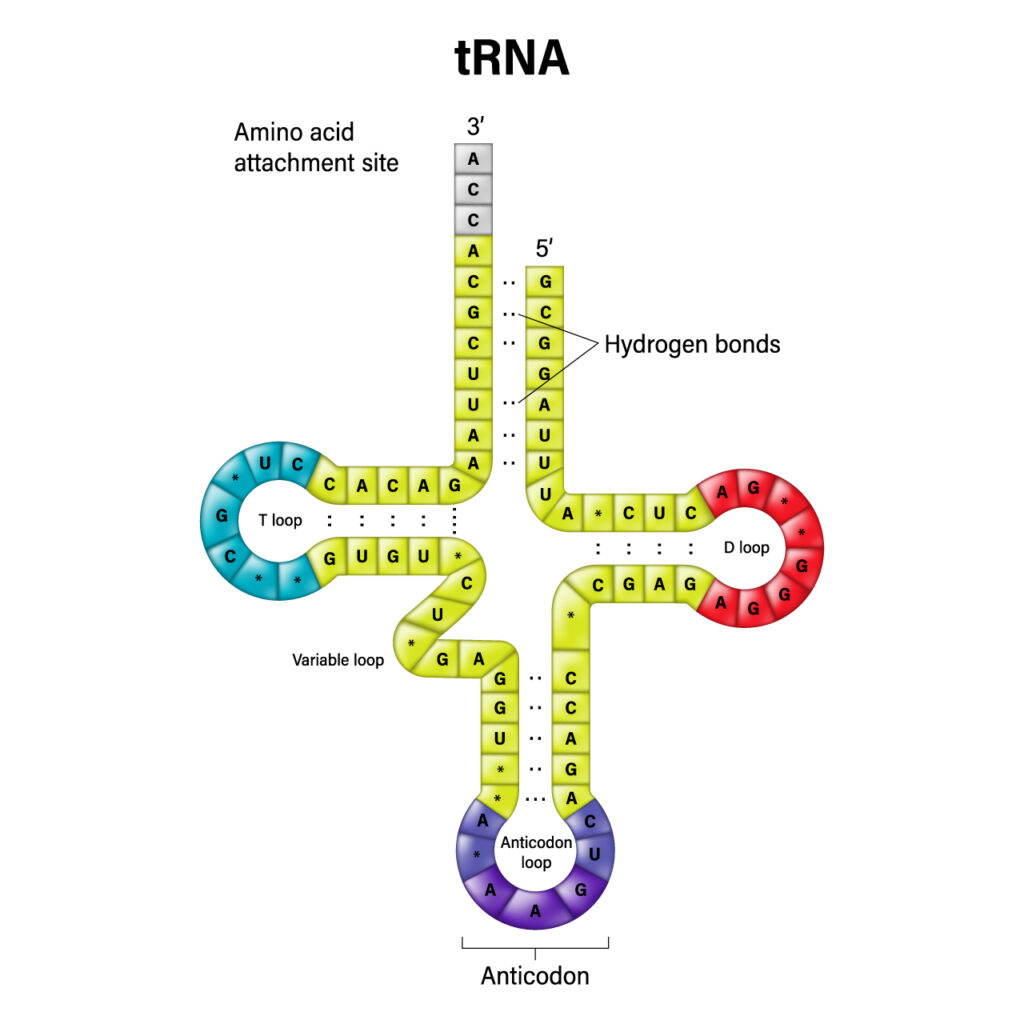
As mRNA is translated in the ribosome, tRNA is the molecule whose anticodon region forms base pairs with the complementary sequence on the mRNA. In doing so, its amino acid is transferred to the ribosome and attached to the growing polypeptide chain to form the protein. This process continues until the ribosome reaches a termination codon on the mRNA, which signals that the protein is complete. In a perfect world, the process of protein synthesis would result in the formation of perfectly functioning proteins in optimal amounts for a healthy, thriving human.
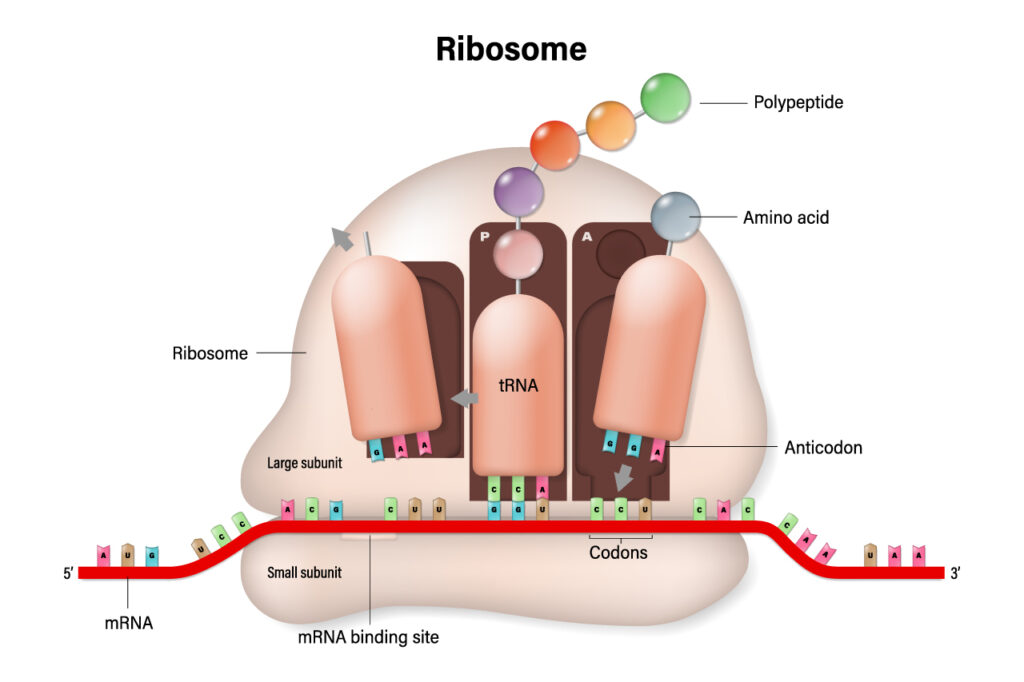
As we know, this often is not always the case. Genetic mutations can cause the correct codon to change to another which results in an incorrect amino acid being added to the protein. An incorrect sequence can also result in a premature termination codon (PTC).
Only 3 of the 64 codons signal termination of the translation. These three termination codons are UAA, UAG, and UGA (1, 2). However, a nonsense mutation switches one nucleotide in an amino acid codon to a different nucleotide that now creates a premature termination codon.
When this happens, the cell does not make the fully formed protein. The resulting truncated protein will either be non-functional or actively harmful. Either case can result in dysfunction and disease. In fact, nonsense mutations cause nearly 1,000 serious genetic disorders (1).
Hence, the multitude of scientists striving to understand the intricacies of the human body and protein synthesis, and how to leverage the knowledge to create solutions to treat disease. As we understand more about how the disease process begins inside the cell, more possible solutions occur. tRNA is showing promise, with one particular therapeutic application in correcting PTCs caused by nonsense mutations.
Potential as a Therapeutic Modality
This is not a new idea. In fact, 40 years ago Yuet Wai Kan’s laboratory at the University of California, San Francisco published a paper on using tRNA to restore protein production (3). Other groups have also validated the approach over the years (4). Research in tRNA has not been widely pursued but some scientists have continued researching it. Recently, it has shown promising results and companies are now investing in the development of tRNA technologies.
Many have focused on suppressor tRNAs (sup-tRNAs). The goal of suppressor tRNA is to override the premature termination codon (PTC), caused by a nonsense mutation, and deliver the amino acid that should be next in the chain. Nucleotides in the anticodon can be changed to make a synthetic tRNA that will bind to a PTC, which is relatively simple to do. Suppressor tRNAs should restore protein production from faulty genes.
Christopher Ahern and John Lueck built a library of hundreds of anticodon-edited suppressor tRNAs (ACE-tRNAs) that can suppress premature termination codons and incorporate desired amino acids (5) which is being utilized to develop therapeutic applications.
Over 10% of all genetic diseases are caused by premature termination codons. Since many different diseases are caused by the same PTC, it is thought that a single suppressor tRNA therapy created for a particular termination codon could potentially work for any disease caused by that same PTC.
Enhancer tRNAs are another type of therapy that has been designed by Tevard Biosciences. Enhancer tRNAs increase production of protein in people who only make half the necessary amount due to haploinsufficiency.
Additionally, the ability to control tRNA provides the ability to also control which mRNAs get translated. In this way, it may be possible to control biological processes that ultimately result in disease (6).
tRNA is also involved in cell division, and down-regulating tRNA could possibly be a way to slow cancer growth (7). Researchers have also discovered that tRNA has many other functions such as gene expression, protein degradation, and apoptosis (8, 9, 10).
Of course, as research continues and knowledge grows, other ways to treat or prevent disease through the use of tRNAs as clinical therapeutics may be discovered.
Challenges and Advantages of Engineered tRNA
Safety is a concern with tRNA therapeutics, especially whether it will cause autoimmunity, off-target activity, and toxicity. Will tRNA bind only with the premature termination codons, or will bind to normal termination codons (1)? If suppressor tRNA binds to the normal termination codons, other proteins will not end where they should, which could create toxic proteins and would likely have severe unintended effects. Mysteriously, this does not seem to be the case, and Ahern, Lueck, and scientists at the Cystic Fibrosis Foundation are not quite sure why their molecules are not causing many more termination codons to be ignored than is normal at this point. Additionally, Zoya Ignatova at the University of Hamburg has engineered tRNA to preferentially bind to premature termination codons, which mitigates this concern.
tRNA faces similar delivery challenges to other nucleic acid therapies, such as being easily degraded, and it must be able to reach the desired cell type and permeate the cell membrane. However, tRNA does have the advantage of a smaller size, and so is easier to deliver through multiple delivery options. It is believed that tRNA will be able to be delivered utilizing technologies that are effective in delivering other oligonucleotide and gene therapies (1). These include lipid nanoparticles (LNP) containing synthetic tRNA or through engineering a virus, in this case an adeno-associated virus (AAV) with DNA to create suppressor tRNA once in the cell.
Another question that must be answered is how many tRNAs must be delivered to gain therapeutic effect? The therapeutic threshold often differs between different diseases, among individuals with the same disease, as well as in different tissues in a single individual.
If you would like to read an in-depth explanation of tRNA biogenesis, processing, interactions, and therapeutic promise and challenges, see this article.
Creating tRNA therapeutics
At the University of Iowa, Christopher Ahern, PhD and his team, along with co-investigator Aislinn Williams, MD, PhD, are using engineered suppressor tRNAs to treat a form of autism associated with loss of function mutations in the SCN2A gene that causes reduced expression of the Nav1.2 sodium channel (11). The tRNAs will be delivered via adeno-associated virus into induced pluripotent stem cell-derived cortical neurons from individuals with nonsense mutations in SCNA2. This tRNA therapy is intended to repair premature termination codons with the goal of restoring Nav1.2 channel function and neurophysiology.
Alltrna has been developing a broad tRNA platform to explore tRNA biology. Alltrna is about “leveraging and understanding the totality of the biology and using the preciseness of this molecule to treat millions and millions of patients (6).” They have built proprietary tools to carry out basic methodologies for expressing, quantifying, modifying, and synthesizing tRNAs to “learn the rules of tRNA as a programmable medicine.”
tRNA can be modified to modulate activity, stability, selectivity, and delivery. Alltrna is learning how to modulate how many of each tRNA are in a cell to change the amount and type of protein in a cell.
Another interesting aspect that they are exploring involves tRNA fragments that function in and in between cells. These fragments have many different novel functions outside of decoding, some of which link to cell proliferation and cancer metastasis.
Tevard Biosciences is developing both suppressor and enhancer tRNA therapies, using AAVs for delivery which would potentially allow for a single dose with permanent results. Both approaches are highly selective and restore gene expression to normal levels, regardless of the position of the mutation in the gene.
Their lead CNS indication is Dravet Syndrome. When caused by heterozygous loss of function mutations in the sodium channel SCN1A, Dravet Syndrome could be treated with a cocktail of three enhancer tRNAs that could double productivity of the working copy of SCN1A in affected cells. This could potentially provide treatment for a broad range of patients with Dravet, no matter what specific defect is in the other copy of SCN1A. However, approximately 25% of people with Dravet also have premature stop mutations in the gene, allowing for use of suppressor tRNA, as well.
Thus, they are exploring a dual approach, in partnership with Zogenix, a company that specializes in epilepsy drugs, to advance both enhancer tRNA and suppressor tRNA to treat Dravet syndrome. Tevard is also developing cures for other genetic epilepsies and both CNS and non-CNS indications caused by haploinsufficiency and/or nonsense mutations.
hC Bioscience is also working on two different, complementary platforms. PTCH (Patch) employs suppressor tRNAs to override PTCs and restore protein function. The second tRNA-based platform is called SWTX (“Switch”) which targets and corrects missense mutations that cause disease. Christopher Ahern, whose work was previously mentioned, founded the company with Leslie J. Williams to translate and augment work performed in the Ahern lab with the goal of fighting cancer and rare diseases.
Dr. Zoya Ignatova who leads a team at the University of Hamburg in Germany is co-developing suppressor tRNA drugs with Arcturus Therapeutics, a company that has developed multiple mRNA medicines (4). Dr. Ignatova, Dr. Suki Albers, and others on their team have experimented with changing parts of the structure of the tRNA molecule and found that it enhances suppression activity (12).
QiXia Decode Therapeutics is a startup founded by Qing Xia of Peking University. QiXia Decode Therapeutics is developing tRNA-enzyme pairs to treat muscular dystrophies and cancers caused by nonsense mutations (4, 13).
ReCode Therapeutics’ proprietary LNP delivery platform called SORT can carry different types of cargo, including tRNA. LNP delivery was chosen because it allows for better control of the drug with the ability to stop, space, and adjust doses. Recode’s lead programs are working on treatments for cystic fibrosis (CF) caused by nonsense mutations and primary ciliary dyskinesia (PCD).
Shape Therapeutics has developed multiple proprietary RNA technologies. The company is pursuing suppressor tRNA to treat Rett syndrome caused by nonsense mutations in the MECP2 gene (4).
In addition to the diseases mentioned, scientists are working on developing tRNAs to treat additional neurodegenerative diseases, psychiatric disorders, genetic forms of blindness, and Hurler Syndrome (a lysosomal storage disease).
Whether tRNA becomes used solely as a stand-alone treatment for a specific mutation, in conjunction with other RNA or gene-editing technologies, or as a broad treatment for any disease with a specific premature termination codon, it has the potential to restore health to thousands and even millions of people. It will be fascinating to watch as tRNA is deeply studied and more tRNA therapies are developed.
References:
- Porter JJ, Heil CS, Lueck JD. Therapeutic promise of engineered nonsense suppressor tRNAs. WIREs RNA. 2021;12:e1641. doi.org/10.1002/wrna.1641
- Katoh T, Iwane Y, Suga H. tRNA engineering for manipulating genetic code. RNA Biol. 2018;15(4-5):453-460. doi: 10.1080/15476286.2017.1343227. Epub 2017 Sep 6. PMID: 28722545; PMCID: PMC6103704.
- Temple GF, Dozy AM, Roy KL, Kan YW. Construction of a functional human suppressor tRNA gene: an approach to gene therapy for beta-thalassaemia. Nature. 1982 Apr 8;296(5857):537-40. doi: 10.1038/296537a0. PMID: 6803169.
- Dolgin E. tRNA therapeutics burst onto startup scene. Nat Biotechnol. 2022 Feb 24. doi: 10.1038/s41587-022-01252-y. Epub ahead of print. PMID: 35210613.
- Lueck JD, Yoon JS, Perales-Puchalt A, Mackey AL, Infield DT, Behlke MA, Pope MR, Weiner DB, Skach WR, McCray PB Jr, Ahern CA. Engineered transfer RNAs for suppression of premature termination codons. Nat Commun. 2019 Feb 18;10(1):822. doi: 10.1038/s41467-019-08329-4. PMID: 30778053; PMCID: PMC6379413.
- Biotech 2050, 2022. Episode 91. [podcast] Lovisa Afzelius, Founding CEO & Theonie Anastassiadis, Co-founder and CIO | Alltrna. Available at: <https://soundcloud.com/user-306573075/91-trna-therapies-lovisa-afzelius-founding-ceo-theonie-anastassiadis-co-founder-cio-alltrna> [Accessed 18 March 2022].
- Aharon-Hefetz N, Frumkin I, Mayshar Y, Dahan O, Pilpel Y, Rak R. Manipulation of the human tRNA pool reveals distinct tRNA sets that act in cellular proliferation or cell cycle arrest. Elife. 2020 Dec 24;9:e58461. doi: 10.7554/eLife.58461. PMID: 33357381; PMCID: PMC7781600.
- Kirchner, S., & Ignatova, Z. (2015). Emerging roles of tRNA in adaptive translation, signalling dynamics and disease. Nature Reviews. Genetics, 16(2), 98– 112. doi.org/10.1038/nrg3861
- Raina, M., & Ibba, M. (2014). tRNAs as regulators of biological processes. Frontiers in Genetics, 5, 171. doi.org/10.3389/fgene.2014.00171
- Schimmel, P. (2018). The emerging complexity of the tRNA world: Mammalian tRNAs beyond protein synthesis. Nature Reviews. Molecular Cell Biology, 19(1), 45– 58. doi.org/10.1038/nrm.2017.77
- Kruth KA, Grisolano TM, Ahern CA, Williams AJ. SCN2A channelopathies in the autism spectrum of neuropsychiatric disorders: a role for pluripotent stem cells? Mol Autism. 2020 Apr 7;11(1):23. doi: 10.1186/s13229-020-00330-9. PMID: 32264956; PMCID: PMC7140374.
- Albers S, Beckert B, Matthies MC, Mandava CS, Schuster R, Seuring C, Riedner M, Sanyal S, Torda AE, Wilson DN, Ignatova Z. Repurposing tRNAs for nonsense suppression. Nat Commun. 2021 Jun 22;12(1):3850. doi: 10.1038/s41467-021-24076-x. PMID: 34158503; PMCID: PMC8219837.
- Shi, N., Yang, Q., Zhang, H. et al. Restoration of dystrophin expression in mice by suppressing a nonsense mutation through the incorporation of unnatural amino acids. Nat Biomed Eng 6, 195–206 (2022). doi.org/10.1038/s41551-021-00774-1
Disclaimer:
“The views, opinions, findings, and conclusions or recommendations expressed in these articles and highlights are strictly those of the author(s) and do not necessarily reflect the views of the Oligonucleotide Therapeutics Society (OTS). OTS takes no responsibility for any errors or omissions in, or for the correctness of, the information contained in these articles. The content of these articles is for the sole purpose of being informative. The content is not and should not be used or relied upon as medical, legal, financial, or other advice. Nothing contained on OTS websites or published articles/highlights is intended by OTS or its employees, affiliates, or information providers to be instructional for medical diagnosis or treatment. It should not be used in place of a visit, call, consultation, or the advice of your physician or other qualified health care provider. Always seek the advice of your physician or qualified health care provider promptly if you have any healthcare-related questions. You should never disregard medical advice or delay in seeking it because of something you have read on OTS or an affiliated site.”