
Imaging small molecule-induced endosomal escape of siRNA
Du Rietz H, Hedlund H, Wilhelmson S, Nordenfelt P, Wittrup A.
Joint First Authors

Hampus Du Rietz and Dr. Anders Wittrup
Delivery is currently the biggest hurdle oligonucleotide therapeutics face to become effective treatments. Oligonucleotide therapeutics not only need to reach the correct tissue and cell type within that tissue, they also have to enter the target cells and subsequently escape the endosomal compartment to be functional. To date, targeted delivery to a particular cell type has been solved with the use of GalNAc conjugation (read more here) while intrathecal delivery allows more unspecific targeting of brain tissue in general (e.g. nusinersen for the treatment of SMA). Yet, even in these cases the vast majority of the therapeutic oligonucleotide remains trapped in the endosomal compartment. It is still unclear how the small functional amount escapes.
To learn more about endosomal escape, watch Prof. Steven Dowdy’s webinar on the subject:
Here, Du Rietz and colleagues from Lund University use live-cell microscopy to study release of cholesterol-conjugated siRNAs from endosomes. They first identified galectin-9 as superior to the previously established galectin-3 membrane damage sensor. They then showed that endolysosomal membrane damage in response to the cationic amphiphilic drugs chloroquine, siramesine and amitriptyline increased siRNA-mediated knockdown dependent on dose and time. Knockdown was positively associated with increases in the number of galectin-9 foci within cells and significantly higher with chloroquine than siramesine or amitriptyline.
To clarify the reasons for the difference between the drugs, the authors studied the redistribution of dextran from endolysosomal compartments to the cytoplasm after treatment with these drugs. They found that chloroquine induces increased dextran release compared to siramesine due to both a higher rate of targeting dextran containing vesicles and higher dextran content in the targeted vesicles. Similar results were seen for release of cholesterol-conjugated siRNA, with the increased efficacy of chloroquine due to targeting of siRNA containing vesicles at a higher rate. Together with results from other endosomal markers, these results strongly suggest that these drugs disrupt different subsets of endolysosomal compartments.
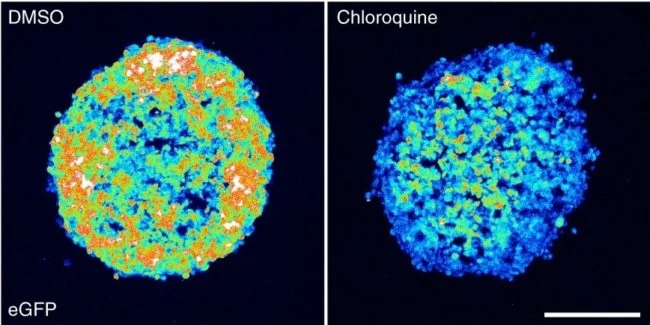
Further experiments demonstrated that using small molecule disruption of endosomal membranes increased siRNA efficacy in other cell lines, but the effect of the drugs varied between cell lines. Membrane disruption using chloroquine or loperamide also improved efficacy and distribution of chol-siRNA into the core of cancer cell spheroids as shown by accumulation of galectin-9 foci.
Live-cell microscopy in combination with the membrane-damage sensor galectin-9 enabled an in-depth study of siRNA release from the endolysosomal compartment.
Why you should read it
Increased understanding of siRNA endosomal release and how to better achieve it will significantly improve the therapeutic index of oligonucleotide therapeutics.